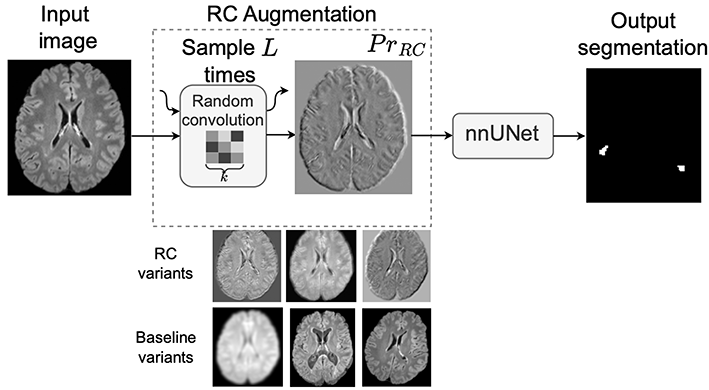
Figure 1: Source: [2]
AI and deep learning have recently transformed medical imaging by enabling automated analysis of complex radiological data, such as detecting lesions, segmenting organs, and predicting disease progression. These methods learn visual representations directly from large datasets and have achieved impressive results across many clinical tasks.
In standard computer vision tasks, deep learning models often suffer from overfitting, where they learn dataset-specific patterns instead of generalizable visual concepts. For example, an object recognition model trained only on images taken with a specific camera or under certain lighting conditions may perform poorly when evaluated on images from a different camera, environment, or background. Although the objects remain the same, subtle changes in image statistics can cause a significant drop in performance.
A similar problem occurs in AI for medical imaging, where domain shift arises when training and test data differ due to variations in scanner vendors, magnetic field strengths, acquisition protocols, or reconstruction pipelines. In clinical tasks such as multiple sclerosis (MS) lesion segmentation, these differences can substantially alter image appearance, leading models to fail when deployed across institutions or imaging centers.
Standard data augmentations like rotations, flips, and intensity perturbations help to some extent, but they often fail to capture realistic variations in imaging characteristics. Random Convolutions (RC) offer a simple yet powerful alternative by exposing models to a wide range of plausible texture and intensity variations while preserving the global structure of anatomical objects [1, 2]. In this blog article, we present our recent work on RC in medical imaging and discuss its practical implications in detail [2].
What Are Random Convolutions and How Are They Used in Practice?
As shown in Figure 1, A convolutional neural network (CNN) is a type of neural network commonly used for image analysis that processes images using small filters (called convolutions) to detect patterns such as edges, textures, and shapes. Random Convolutions (RC) augment an input image by passing it through a shallow convolutional network with randomly sampled filters. The network has depth L, where depth refers to the number of convolutional layers in the CNN, and each convolution uses random kernels of spatial size k × k.
The input image can be viewed as a tensor of size C × H × W, where C denotes the number of channels and H and W denote the spatial dimensions. Each random convolution layer applies a kernel that spans all C channels and has spatial size k × k. The kernel values are randomly sampled from a zero-mean Gaussian distribution with variance σ², and a new set of kernels is drawn for each layer ℓ = 1, …, L.
The RC transformation is obtained by sequentially applying the L random convolution layers to the input image, resulting in a transformed image that preserves the overall anatomical structure but exhibits altered texture statistics. The kernels are re-sampled for each mini-batch, gradients are not propagated through them, and only the parameters of the downstream segmentation network are optimized.
In practice, RC is used as a preprocessing augmentation. The input image is first transformed by the random convolution module and then processed by a standard segmentation framework. At inference time, the RC module is removed and the network operates on the original, unaltered images. Empirically, RC preserves anatomical structures while introducing significant textural variation, improving robustness to domain shift without degrading in-domain performance.
Random Convolutions for Domain Randomization and Improved Generalization
RC can be viewed as a form of implicit domain randomization. Instead of explicitly modeling scanner physics or performing computationally expensive style transfer, RC generates new domains by perturbing texture statistics in a simple and task-agnostic manner. This highlights an important insight: robustness does not require realistic simulations of new domains—often, it is sufficient to prevent the model from relying on spurious appearance cues.
Convolutional neural networks are known to rely heavily on local texture cues. Although this inductive bias can improve performance on in-distribution data, it becomes problematic under domain shift, where texture statistics differ between training and deployment data, as is common in medical imaging. By applying L random convolutions with kernel size k, RC introduces strong textural variability while preserving underlying anatomical structure. This encourages the network to rely more on shape, spatial context, and global morphology, which tend to be more invariant to scanner and protocol variability.
Limitations and Extensions
The effectiveness of RC depends on the choice of depth L and kernel size k. Very large values can distort semantic content, while overly small values may not introduce sufficient variability.
To address this, later work proposed progressive random convolutions, which carefully control the depth and strength of random filtering to balance appearance diversity and structural fidelity.
Key Takeaways
- Random convolutions use a shallow network of depth L with random kernels of size k as a data augmentation strategy.
- They introduce strong textural variability while preserving anatomical structure.
- In medical imaging tasks such as MS lesion segmentation, they significantly improve robustness to domain shift.
- Their simplicity and low computational cost make them well suited for real-world deployment with heterogeneous data.
Random convolutions show that carefully controlled randomness at the input level can substantially improve how models generalize beyond the data they were trained on.
References
[1] Choi, S., Das, D., Choi, S., Yang, S., Park, H., & Yun, S. (2023).
Progressive Random Convolutions for Single Domain Generalization.
Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 10312–10322.
[2] Varma, A., Scholz, D., Erdur, A. C., Peeken, J. C., Rueckert, D., and Wiestler, B. (2026).
Improving out-of-domain generalization in Multiple Sclerosis detection and segmentation using Random Convolutions.
Pattern Recognition Letters, Elsevier (ScienceDirect).